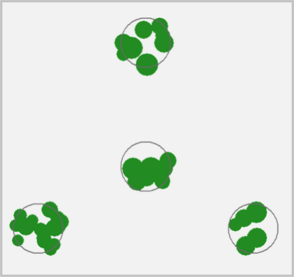
Predict plot-level canopy cover from individual tree measurements
Source:R/calc_tcc_metrics.R
calc_tcc_metrics.Rdcalc_tcc_metrics() computes predicted plot-level tree canopy cover (TCC)
from standard field inventory measurements. By default, the output includes
a full set of stand structure variables used to model the plot-level TCC
value (see Details).
Arguments
- tree_list
A data frame with tree records for one FIA plot. In general, the input data frame will have the columns specified in DEFAULT_TREE_COLUMNS (see
?DEFAULT_TREE_COLUMNS). Potentially, only a subset of those columns will be needed depending on values given for the argumentsstem_mapandfull_outputdescribed below. If the input data frame has a column named"CRWIDTH"it will be used for tree crown width values, otherwise, crown widths will be calculated with a call tocalc_crwidth().- stem_map
A logical value indicating whether to map individual tree stems explicitly, using coordinates specified in terms of distance and azimuth from subplot/microplot centers. The default is
TRUE, in which case the inputtree_listmust contain columns"DIST"and"AZIMUTH". This argument may be set toFALSEif individual tree locations are not available, in which case TCC will be predicted assuming a random arrangement of tree locations (see Details).- full_output
A logical value indicating whether to include the full set of components used to derive the plot-level prediction. By default, the output list includes subplot-level TCC estimates, live tree and sapling counts, stand height metrics, and point pattern statistics, depending on the value given for
stem_map(see Details).- digits
Optional integer indicating the number of digits to keep in the return values (defaults to
1). May be passed tocalc_crwidth()andcalc_ht_metrics().- ...
Optional arguments passed to
create_fia_ppp()ifstem_map = TRUE.
Value
If full_output = TRUE, a named list with element model_tcc containing
the plot-level predicted tree canopy cover as percent (0:100), and
additional named elements containing stand structure metrics as described in
Details. If full_output = FALSE, a single numeric value of plot-level
predicted TCC is returned instead.
Details
This function provides two methods for predicting plot-level TCC.
The default "stem-map" method requires individual tree coordinates to be
given in the input as distance and azimuth from subplot centers for trees
with diameter >= 5 in. (12.7 cm), and from microplot centers for
"saplings" having diameter >= 1 in. (2.54 cm) but < 5 in.
(12.7 cm). This method involves mapping trees spatially within the plot
boundary to account for crown overlap explicitly, along with empirical
modeling of the understory sapling contribution to total canopy cover
(Toney et al. 2009). The empirical model for the sapling component also uses
the spatial point pattern of overstory trees as a predictor variable (using
a square root transformation of Ripley's edge-corrected K-function, Ripley
1977, Stoyan and Penttinen 2000).
Alternatively, TCC can be predicted using a simplified approach that does not
include exact stem placement within the plot boundary (stem_map = FALSE). A
random arrangement of stems is assumed in that case. This is the method used
to estimate tree canopy cover in the Forest Vegetation Simulator (Crookston
and Stage 1999).
Both methods require estimates of individual tree crown widths, which are
computed with calc_crwidth() if not provided in the input tree list.
The stem-map method also requires computation of several stand structure metrics which are used in various components of the overall model used to derive a plot-level TCC estimate. These additional variables include:
individual subplot and microplot crown overlays via
calc_crown_overlay()a stand height metric (
meanTreeHtBAW), and plot-level counts of mature trees and saplings, viacalc_ht_metrics()descriptive spatial statistics for the overstory tree point pattern via
create_fia_ppp() |> spatstat.explore::Lest()
By default, calc_tcc_metrics() returns a named list containing the
plot-level modeled TCC value, along with those additional component
variables. Specific elements of the returned list include some or all of the
following, conditionally:
$model_tcc: plot-level predicted canopy cover of trees>= 1inch (2.54cm) diameter, derived by one of the two methods described above depending on the value given for argumentstem_map = TRUE|FALSE
If the stem-map method is used, then TCC values derived by crown overlay on the individual subplot and microplot boundaries are included, along with means of the four subplot/microplot values:
$subpN_crown_overlay: estimated canopy cover of trees>= 5-in.(12.7 cm) diameter in subplotNbased on crown overlay (N = 1:4)$subp_overlay_mean: mean of the four subplot crown overlays$micrN_crown_overlay: estimated canopy cover of saplings in the microplot of subplotNbased on crown overlay (N = 1:4)$micr_overlay_mean: mean of the four microplot crown overlays
A set of spatial point pattern statistics is also included when the stem-map
method is used. A square root transformation of Ripley's K function using
isotropic edge correction is computed with spatstat.explore::Lest() for
trees >= 5-in. (12.7 cm) diameter within the four-subplot observation
window. The mean of the following values is a predictor variable in a linear
regression model used to estimate the sapling contribution to total tree
canopy cover:
$L_rft: estimates of the L-function atrfeet (r=6,8,10, and12)
If the argument full_output = TRUE (the default), then the output will
also include all of the the named elements from the output of
calc_ht_metrics().
References
Crookston, N.L. and A.R. Stage. (1999). Percent canopy cover and stand structure statistics from the Forest Vegetation Simulator. Gen. Tech. Rep. RMRS-GTR-24. Ogden, UT: U. S. Department of Agriculture, Forest Service, Rocky Mountain Research Station. 11 p. https://research.fs.usda.gov/treesearch/6261.
Ripley, B.D. (1977). Modelling spatial patterns. Journal of the Royal Statistical Society: Series B (Methodological), 39(2): 172–192. https://doi.org/10.1111/j.2517-6161.1977.tb01615.x.
Stoyan, D., and Penttinen, A. (2000). Recent applications of point process methods in forestry statistics. Statistical Science, 15(1), 61–78. http://www.jstor.org/stable/2676677.
Toney, C., J.D. Shaw and M.D. Nelson. 2009. A stem-map model for predicting tree canopy cover of Forest Inventory and Analysis (FIA) plots. In: McWilliams, Will; Moisen, Gretchen; Czaplewski, Ray, comps. Forest Inventory and Analysis (FIA) Symposium 2008; October 21-23, 2008; Park City, UT. Proc. RMRS-P-56CD. Fort Collins, CO: U.S. Department of Agriculture, Forest Service, Rocky Mountain Research Station. 19 p. https://research.fs.usda.gov/treesearch/33381.
Examples
# using the spatially explicit "stem-map model" by default
calc_tcc_metrics(plantation)
#> $model_tcc
#> [1] 88.4
#>
#> $subp1_crown_overlay
#> [1] 86.8
#>
#> $subp2_crown_overlay
#> [1] 91.7
#>
#> $subp3_crown_overlay
#> [1] 80.2
#>
#> $subp4_crown_overlay
#> [1] 87.2
#>
#> $subp_overlay_mean
#> [1] 86.475
#>
#> $micr1_crown_overlay
#> [1] 0
#>
#> $micr2_crown_overlay
#> [1] 0
#>
#> $micr3_crown_overlay
#> [1] 20.2
#>
#> $micr4_crown_overlay
#> [1] 22.5
#>
#> $micr_overlay_mean
#> [1] 10.675
#>
#> $L_6ft
#> [1] 3.868305
#>
#> $L_8ft
#> [1] 6.627377
#>
#> $L_10ft
#> [1] 7.300455
#>
#> $L_12ft
#> [1] 11.35045
#>
#> $numTrees
#> [1] 89
#>
#> $meanTreeHt
#> [1] 44.8
#>
#> $meanTreeHtBAW
#> [1] 45.3
#>
#> $meanTreeHtDom
#> [1] 44.8
#>
#> $meanTreeHtDomBAW
#> [1] 45.3
#>
#> $maxTreeHt
#> [1] 51
#>
#> $predomTreeHt
#> [1] 50.7
#>
#> $numSaplings
#> [1] 2
#>
#> $meanSapHt
#> [1] 34.5
#>
#> $maxSapHt
#> [1] 43
#>
# return only the predicted TCC value (`$model_tcc`)
calc_tcc_metrics(plantation, full_output = FALSE)
#> [1] 88.4
# using the "FVS method" which assumes random tree locations
calc_tcc_metrics(plantation, stem_map = FALSE, full_output = FALSE)
#> [1] 81.4